This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Subcellular localization
The localization of Notch1 is predicted using the following databases:
Database: LOCATE
Prediction: plasma membrane, nucleus
Database: Hum-mPLoc, Euk-mPLoc
Prediction: nucleus
Database: SherLoc2
Prediction: plasma membrane (0.93), lysosomal (0.04), extracellular (0.01), Golgi apparatus (0.01)
Database: LOCATE
Prediction: plasma membrane, nucleus
Database: Hum-mPLoc, Euk-mPLoc
Prediction: nucleus
Database: SherLoc2
Prediction: plasma membrane (0.93), lysosomal (0.04), extracellular (0.01), Golgi apparatus (0.01)
|
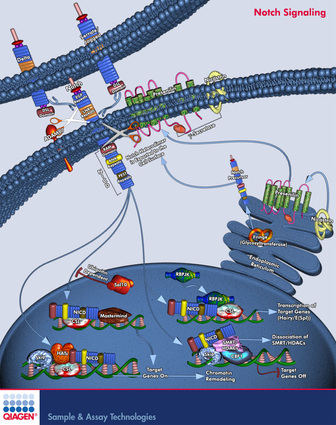
_Figure 1: Figure showing subcellular localization of Notch1 preprotein and processed form. Dark blue structure at the bottom of the figure represents nucleus. Lipid bilayer of plasma membrane is represented by the dark blue bubble-like structure. The top cell represents signal transmission cell, and the lower cell represents the signal receiving cell. [Figure from https://www.qiagen.com/geneglobe/pathwayview.aspx?pathwayID=325]
|
Analysis
The predicted localization of Notch1 is consistent with its function in Notch signalling pathway and transcriptional regulation [UniProt]. It is known that Notch1preprotein is cleaved upon binding of its ligand. One fragment of the protein translocates to the nucleus and regulates the transcription of genes. Hence, it is not surprising that the protein could be found in plasma membrane (as receptors) and nucleus (in its cleaved form). However, SherLoc2 prediction did not report localization of Notch1 to the nucleus. This is perhaps because the entire Notch1 preprotein sequence was used to generate the prediction. Indeed, when the truncated Notch1 protein sequence was entered into SherLoc2, the program reported localization of Notch1 in nucleus.
The predicted localization of Notch1 is consistent with its function in Notch signalling pathway and transcriptional regulation [UniProt]. It is known that Notch1preprotein is cleaved upon binding of its ligand. One fragment of the protein translocates to the nucleus and regulates the transcription of genes. Hence, it is not surprising that the protein could be found in plasma membrane (as receptors) and nucleus (in its cleaved form). However, SherLoc2 prediction did not report localization of Notch1 to the nucleus. This is perhaps because the entire Notch1 preprotein sequence was used to generate the prediction. Indeed, when the truncated Notch1 protein sequence was entered into SherLoc2, the program reported localization of Notch1 in nucleus.