This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Post-translational modifications
Many proteins are modified with additional chemical groups or structural changes after being translated by ribosome. In the process of maturation and activation, Notch1 protein is cleaved 3 times [Figure 1]. The first cleavage occurs within its biosynthesis, where a Furin-like protease in the Golgi body cleaves the nascent Notch proteins and converts them into heterodimers to enable their expression at cell surface. After successfully being expressed at the cell surface, Notch1 heterodimer can now play its role as receptors in signaling pathways. When the ligand binds to Notch1 heterodimer, 2 additional cleavages are induced, 1 of which cleaves off most of the Notch1 extracellular domain, whereas the other cleaves the trans-membrane domain and releases the intracellular domain [Qiagen] [1]. The intracellular domain then translocates to the nucleus to regulate gene transcription [1].
Many proteins are modified with additional chemical groups or structural changes after being translated by ribosome. In the process of maturation and activation, Notch1 protein is cleaved 3 times [Figure 1]. The first cleavage occurs within its biosynthesis, where a Furin-like protease in the Golgi body cleaves the nascent Notch proteins and converts them into heterodimers to enable their expression at cell surface. After successfully being expressed at the cell surface, Notch1 heterodimer can now play its role as receptors in signaling pathways. When the ligand binds to Notch1 heterodimer, 2 additional cleavages are induced, 1 of which cleaves off most of the Notch1 extracellular domain, whereas the other cleaves the trans-membrane domain and releases the intracellular domain [Qiagen] [1]. The intracellular domain then translocates to the nucleus to regulate gene transcription [1].
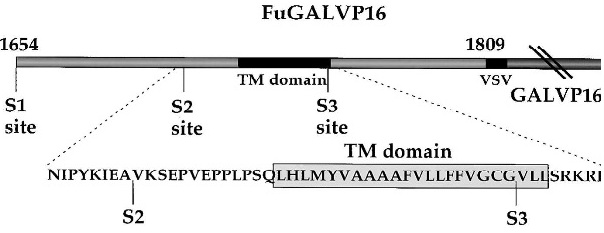
Figure 1: The cleavages are designated S1, S2 and S3 respectively according to order as discussed above. S3 cleavage is within the transmembrane (TM) domain, occurs after binding of ligand, and releases the intracellular domain (right). Extracellular domain is to the left. Figure obtained from [1].
Apart from undergoing post-translational cleavage upon activation, Notch1 is also modified by chemical groups, which are important for the regulation of Notch signaling. The intracellular domain of the cleaved Notch1 can be negatively regulated via ubiquitination by E3-ubiquitin ligase, followed by proteasomal degradation [2]. Apart from that, cyclin-dependent kinase is also reported to negatively regulate the intracellular domain of Notch by phosphorylating its PEST domain [2]. The extracellular domain of Notch could also be post-translationally modified via glycosylation. This affects the affinity Notch receptor to its ligand [3] and thus represent a mechanism for yet another level of Notch signaling regulation.
References:
1. Brou, C. et al. (2000). A Novel Proteolytic Cleavage Involved in Notch Signaling: The Role of the Disintegrin-Metalloprotease TACE. Molecular Cell 5(2): 207-216.
2.Fortini, M.E. (2009). Notch Signaling: The Core Pathway and Its Posttranslational Regulation. Developmental Cell 16: 633-647.
3. Miele, L. (2006). Notch Signaling. Clinical Cancer Research 12: 1074-1079.