DNA motifs
MOTIF identified 14 motifs from the full-length NOTCH1 gene sequence:
|
1. EGF_1 (EGF-like domain signature 1)
Pattern: C-x-C-x(2)-{V}-x(2)-G-{C}-x-C 2. J-ACTX (Janus-faced atracotoxin family signature) Pattern: C-{C}(6)-C-x(2)-C-C-x-C-C-x(4)-C-x(9,10)-C 3. Integrin_Beta (Integrins beta chain cysteine-rich domain signature) Pattern: C-x-[GNQ]-x(1,3)-G-x-C-x-C-x(2)-C-x-C 4. CTCK_1 (C-terminal cystine knot signature) Pattern: C-C-x(13)-C-x(2)-[GN]-x(12)-C-x-C-x(2,4)-C 5. ANAPHYLATOXIN_1 (Anaphylatoxin domain signature) Pattern: [CSH]-C-x(2)-[GAP]-x(7,8)-[GASTDEQR]-C-[GASTDEQL]- x(3,9)-[GASTDEQN]-x(2)-[CE]-x(6,7)-C-C 6. I_CONOTOXIN (I-superfamily conotoxin signature) Pattern: C-{C}(6)-C-{C}(5)-C-C-x(1,3)-C-C-x(2,4)-C-x(3,10)-C 7. AGOUTI_1 (Agouti domain signature) Pattern: C-x(6)-C-x(6)-C-C-x(2)-C-x(2)-C-x-C-x(6)-C-x-C-x(6,9)-C |
8. IGFBP_N_1 (Insulin-like growth factor-binding protein N-terminal domain signature)
Pattern: [GP]-C-[GSET]-[CE]-[CA]-x(2)-C-[ALP]-x(6)-C 9. THIOLASE_3 (Thiolases active site) Pattern: [AG]-[LIVMA]-[STAGCLIVM]-[STAG]-[LIVMA]-C-{Q}-[AG]-x-[AG]-x-[AG]-x-[SAG] 10. TUBULIN (Tubulin subunits alpha, beta, and gamma signature) Pattern: [SAG]-G-G-T-G-[SA]-G 11. VWFC_1 (VWFC domain signature) Pattern: C-x(2,3)-C-{CG}-C-x(6,14)-C-x(3,4)-C-x(2,10)-C-x(9,16)-C-C-x(2,4)-C 12. 4FE4S_FER_1 (4Fe-4S ferredoxin-type iron-sulfur binding region signature) Pattern: C-x-{P}-C-{C}-x-C-{CP}-x-{C}-C-[PEG] 13. 2FE2S_FER_1 (2Fe-2S ferredoxin-type iron-sulfur binding region signature) Pattern: C-{C}-{C}-[GA]-{C}-C-[GAST]-{CPDEKRHFYW}-C 14. DEFENSIN (Mammalian defensins signature) Pattern: C-x-C-x(3,5)-C-x(7)-G-x-C-x(9)-C-C |
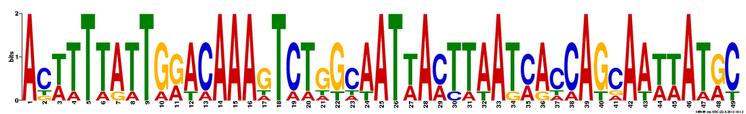
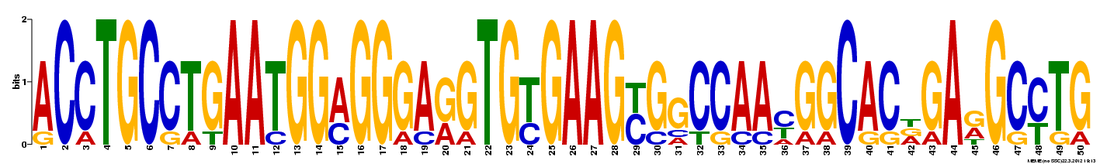
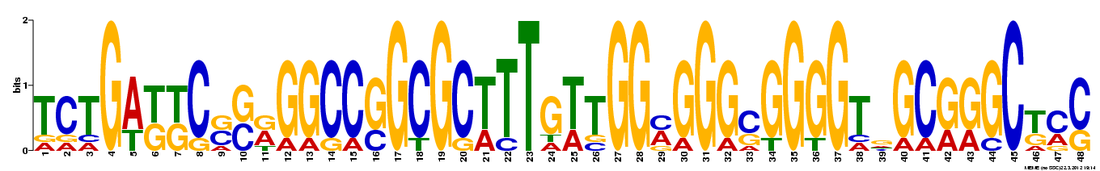
MEME identified 3 conserved DNA motifs for NOTCH1:
Analysis
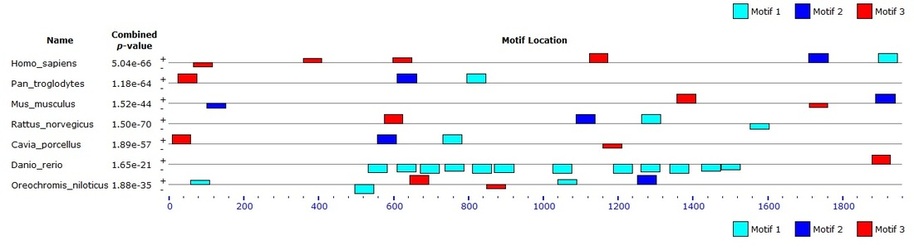
A total of 14 DNA motifs were identified via MOTIF by entering the whole gene sequence of NOTCH1 into the database. These motifs are consistent with the function of NOTCH1. For instance, its Integrin_beta motif is suggestive of its role as a receptor (PROSITE). In order to investigate how the motifs are conserved in other organisms, I performed DNA motif search on the gene homologs discussed in the DNA homolog page using MEME. The search returned the results below:
A total of 14 DNA motifs were identified via MOTIF by entering the whole gene sequence of NOTCH1 into the database. These motifs are consistent with the function of NOTCH1. For instance, its Integrin_beta motif is suggestive of its role as a receptor (PROSITE). In order to investigate how the motifs are conserved in other organisms, I performed DNA motif search on the gene homologs discussed in the DNA homolog page using MEME. The search returned the results below:
This figure from MEME illustrates that the 3 motifs, represented by colored boxes, are well conserved in all the organisms analyzed. Although the location and distribution of the motifs differ from one organism to another, the presence of the motif in the homologous genes suggest that these motifs may have functional significance. However, based on the motif location, these 3 motifs do not seem to be among the 14 motifs identified via MOTIF. It is possible that these conserved motifs may represent significant motifs that are currently uncharacterized.
References:
1. MOTIF
2. Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994
3. MEME
4. PROSITE