You are here: Protein > Protein-protein Interactions
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Protein-protein interactions
Apart from the protein-protein interactions shown above, Notch1 signaling appears to be intertwined with other signaling pathways such as the anti-apoptotic PI3K/Akt pathway, making these signaling pathways likely major contributors of tumor formation. The activated, cleaved intracellular domain of Notch1 (see Post-translational modifications) is reported to inhibit apoptosis via mTOR, a downstream component of the PI3K/Akt signaling pathway [5]. This interaction inhibits the tumor suppressor protein p53, thereby promoting tumor progression [5].
Analysis:
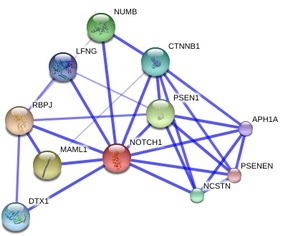
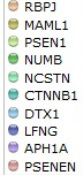
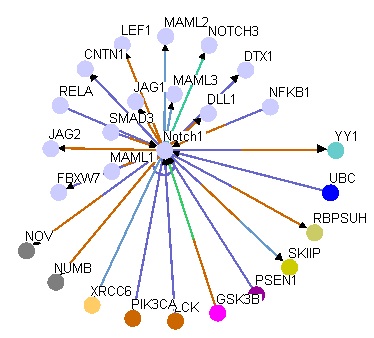
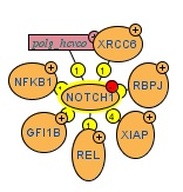
Apart from using the databases above to obtain illustrative figures on the protein-protein interactions of Notch1, I also performed searches using other databases such as BioGRID 3.1, which returned 72 unique protein interactions for Notch1. These matched the results obtained from String 9.0 except for NCSTN and APH1A were detected by String 9.0 but not in BioGRID 3.1. In addition, MINT identified the interaction of Notch1 with XIAP and REL, which were not recognized in other searches. These two protein interactions are especially important in the context of T-ALL because XIAP is an inhibitor of apoptosis, which could lead to lymphoproliferative diseases if misregulated or mutated [MINT] [OMIM], whereas REL is a "proto-oncogene that may play a role in differentiation and lymphopoiesis" [UniProt]. The differences in search results might be due to the large amount of proteins that interact with Notch1, and different databases may use different criteria for scoring the interactions, leading to different results being returned. Personally, I favor String 9.0 over other databases because it provides nice and clear illustration of protein interactions as well as offers user-friendly way to explore related topics, such as experiments and evidences regarding the protein interactions.
The functions of the interacting proteins seem to be consistent with the ontology of Notch1 in regulation of cell fate. For instance, NUMB is reported to play a role in the regulation of cell fate decision during development [String 9.0], DTX-1 "promotes B-cell development at the expense of T-cell development" [UniProt] and MAML1 is a transcriptional co-activator that "controls hematopoietic development by regulating NOTCH-mediated lymphoid cell fate decisions" [UniProt]. From these protein-protein interactions and their functions, it is clear that mis-regulation and/or mutations in Notch1 could lead to aberrancy in T-cell development and consequently contribute to T-ALL.
Apart from using the databases above to obtain illustrative figures on the protein-protein interactions of Notch1, I also performed searches using other databases such as BioGRID 3.1, which returned 72 unique protein interactions for Notch1. These matched the results obtained from String 9.0 except for NCSTN and APH1A were detected by String 9.0 but not in BioGRID 3.1. In addition, MINT identified the interaction of Notch1 with XIAP and REL, which were not recognized in other searches. These two protein interactions are especially important in the context of T-ALL because XIAP is an inhibitor of apoptosis, which could lead to lymphoproliferative diseases if misregulated or mutated [MINT] [OMIM], whereas REL is a "proto-oncogene that may play a role in differentiation and lymphopoiesis" [UniProt]. The differences in search results might be due to the large amount of proteins that interact with Notch1, and different databases may use different criteria for scoring the interactions, leading to different results being returned. Personally, I favor String 9.0 over other databases because it provides nice and clear illustration of protein interactions as well as offers user-friendly way to explore related topics, such as experiments and evidences regarding the protein interactions.
The functions of the interacting proteins seem to be consistent with the ontology of Notch1 in regulation of cell fate. For instance, NUMB is reported to play a role in the regulation of cell fate decision during development [String 9.0], DTX-1 "promotes B-cell development at the expense of T-cell development" [UniProt] and MAML1 is a transcriptional co-activator that "controls hematopoietic development by regulating NOTCH-mediated lymphoid cell fate decisions" [UniProt]. From these protein-protein interactions and their functions, it is clear that mis-regulation and/or mutations in Notch1 could lead to aberrancy in T-cell development and consequently contribute to T-ALL.
References:
1.String 9.0
2.Osprey 1.0.1
3. MINT
4. BioGRID3.1
5. Mungamuri, S.K., Yang, X., Thor, A.D., & Somasundaram, K. (2006). Cancer Research 66(9): 4715-4724. Doi: 10.1158/0008-5472.CAN-05-3830
1.String 9.0
2.Osprey 1.0.1
3. MINT
4. BioGRID3.1
5. Mungamuri, S.K., Yang, X., Thor, A.D., & Somasundaram, K. (2006). Cancer Research 66(9): 4715-4724. Doi: 10.1158/0008-5472.CAN-05-3830