This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Organism phenotypes and RNAi
Organism phenotypes
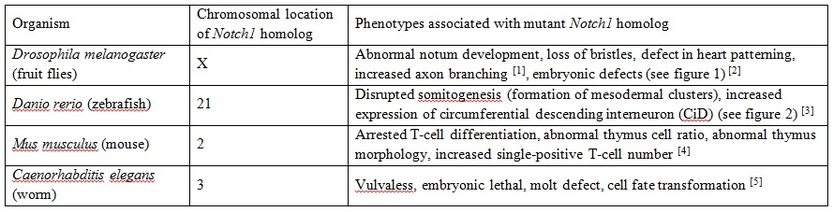
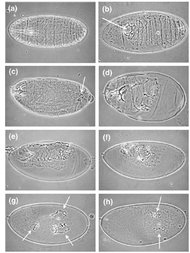
Listed below are some of the phenotypes that are associated with mutations in Notch1 homologs in the corresponding model organism. Phenotypes presented here are selected based on their relatedness to cell fate determination. Abnormalities in cell fate determination could lead to various types of cancers, including T-ALL.
RNAi
Out of the 4 model organisms listed above, I was only successful in obtaining the RNAi information for D.melanogaster and C.elegans. Results are shown below:
D.melanogaster [N]*
- Embryonic defects [2]
- Impaired memory
- Vein thickening
- Increased number of small cells in adult midgut epithelium
C.elegans [lin-12] [5]
- Incomplete vulval morphogenesis
- Multiple anchor cells
- Abnormal vulval cell fate specification
- Embryonic lethal
- Abnormal locomotion
Analysis
Among all the model organisms analyzed above, the mutant phenotypes expressed in mice are the closest manifestations of the human T-ALL. This is not surprising given that mouse is the only mammalian model organism chosen for this analysis. Nevertheless, I was unable to obtain RNAi information in mouse models that are related to T-ALL. This is perhaps because of the difficulties in performing RNAi in mice. As for the other organisms, although they did not express phenotypes that could strongly be associated with T-ALL, mutations of Notch1 homologs in these animals did show abnormalities in cell fate determination, resulting in abnormal cell count and/or abnormal development and function. In short, among these animals, mice may be the most suitable for modeling T-ALL for research purposes. However, for studies of specific aspect of T-ALL and/or Notch signaling, other organisms may be very valuable as well.
Among all the model organisms analyzed above, the mutant phenotypes expressed in mice are the closest manifestations of the human T-ALL. This is not surprising given that mouse is the only mammalian model organism chosen for this analysis. Nevertheless, I was unable to obtain RNAi information in mouse models that are related to T-ALL. This is perhaps because of the difficulties in performing RNAi in mice. As for the other organisms, although they did not express phenotypes that could strongly be associated with T-ALL, mutations of Notch1 homologs in these animals did show abnormalities in cell fate determination, resulting in abnormal cell count and/or abnormal development and function. In short, among these animals, mice may be the most suitable for modeling T-ALL for research purposes. However, for studies of specific aspect of T-ALL and/or Notch signaling, other organisms may be very valuable as well.
References:
1. FlyBase
2. Drosophila pic from Boutla, A., Delidakis, C., Livadaras, l., Tsagris, M., & Tabler, M. (2001). Short 5’-phosphorylated double-stranded RNAs induce RNA interference in Drosophila. Current Biology 11: 1776-1780.
3. ZFIN
4. MGI
5. WormBase
6. Nikolaou, N., Watanabe-Asaka, T., Gerety, S., Distel, M., Köster, R.W., and Wilkinson, D.G. (2009) Lunatic fringe promotes the lateral inhibition of neurogenesis. Development 136(15): 2523-2533.