This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Notch1 protein domain
Domain searches using Pfam returned 40 domains of Notch1, including 34 significant domains and 6 insignificant domains:
To the left of the figure are 29 Epidermal Growth Factor (EGF)-like domains, with the red boxes specifically representing Calcium-binding EGF domains and the blue box representing human growth factor-like EGF. The yellow and purple boxes represent NOD and NODP respectively, both of which belong to the NOTCH protein family and are important for cell differentiation. The 2 orange boxes are the Ankyrin repeats (3 copies each). Lastly, the pink cap at the right end of the figure is a domain with unknown function.
A similar structure is obtained from SMART:
A similar structure is obtained from SMART:
The domains identified were similar, except for the absence of the 3 NL domains in Pfam results and the loss of the NOD and NODP domains in the SMART results. However, based on the descriptions provided for the NL domain, it is likely that NL refers to the NOD and NODP domains in Pfam.
Using MOTIF and selecting the PROSITE pattern database, 5 Notch1 protein motifs were identified: EGF_1, EGF_CA, EGF_2, ASX_HYDROXYL, and WD_REPEATS_1. This result is consistent with the results above. ASX_HYDROXYL (Aspartic acid and asparagine hydroxylation site) is likely to be part of Ankyrin repeats [1], whereas WD_REPEATS_1 (Trp-Asp (WD-40) repeats) may be identical to the domain with unknown function stated above (see below for more information).
Analysis:
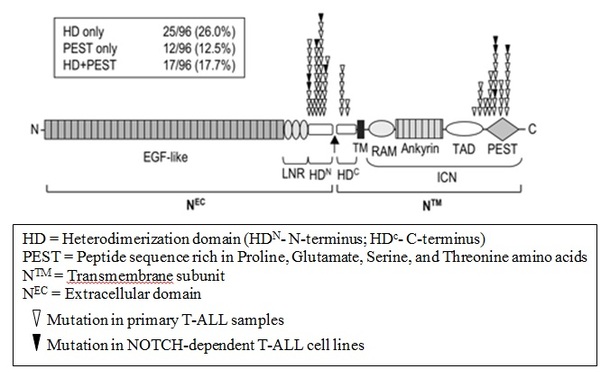
Upon comparing the mutations of Notch1 in T-ALL as cataloged by Weng, P. et al. (2004) [2] and a more recent article by Bhanushali, A.A et al. (2010) [3], it was identified that a significant number of mutations were mapped to the heterodimerization domain (HD) and PEST domain [see figure below]. The HD domain is located in the grey region at the middle of the figure above, where no domains were recognized. As for the PEST domain, published articles stated that it is located at the carboxyl-terminal of Notch1 protein, which would be located at the right end of the figure above where the pink cap is. This is confirmed by looking at the amino acid sequences at the carboxyl terminal of the protein in UniProtKB, where description of the amino acid coordinate is provided. From Pfam, it is noted that the domain with unknown function (pink cap) starts from amino acid 2478 to 2541. This corresponds to the poly-serine region as indicated in UniProtKB, hence indicating that the domain of unknown function is likely to contain the PEST domain, which is characterized by high occurrence of proline (P), glutamate (E), serine (S) and threonine (T).
Using MOTIF and selecting the PROSITE pattern database, 5 Notch1 protein motifs were identified: EGF_1, EGF_CA, EGF_2, ASX_HYDROXYL, and WD_REPEATS_1. This result is consistent with the results above. ASX_HYDROXYL (Aspartic acid and asparagine hydroxylation site) is likely to be part of Ankyrin repeats [1], whereas WD_REPEATS_1 (Trp-Asp (WD-40) repeats) may be identical to the domain with unknown function stated above (see below for more information).
Analysis:
Upon comparing the mutations of Notch1 in T-ALL as cataloged by Weng, P. et al. (2004) [2] and a more recent article by Bhanushali, A.A et al. (2010) [3], it was identified that a significant number of mutations were mapped to the heterodimerization domain (HD) and PEST domain [see figure below]. The HD domain is located in the grey region at the middle of the figure above, where no domains were recognized. As for the PEST domain, published articles stated that it is located at the carboxyl-terminal of Notch1 protein, which would be located at the right end of the figure above where the pink cap is. This is confirmed by looking at the amino acid sequences at the carboxyl terminal of the protein in UniProtKB, where description of the amino acid coordinate is provided. From Pfam, it is noted that the domain with unknown function (pink cap) starts from amino acid 2478 to 2541. This corresponds to the poly-serine region as indicated in UniProtKB, hence indicating that the domain of unknown function is likely to contain the PEST domain, which is characterized by high occurrence of proline (P), glutamate (E), serine (S) and threonine (T).
References:
1. Yang M, Ge W, Chowdhury R, Claridge TD, Kramer HB, Schmierer B, McDonough MA, Gong L, Kessler BM, Ratcliffe PJ, Coleman ML, Schofield CJ. (2011). Asparagine and aspartate hydroxylation of the cytoskeletal ankyrin family is catalyzed by factor-inhibiting hypoxia-inducible factor. J Biol Chem. 286(9):7648-60. Epub 2010 Dec 22. PubMed PMID: 21177872; PubMed Central PMCID: PMC3045019.
2. Weng, A.P., Ferrando, A.A., Lee, W., Morris IV, J.P., Silverman, L.B., Sanchez-Irizarry, C., Blacklow, S.C., Look, A.T., & Aster, J.C. (2004). Activating Mutations of NOTCH1 in Human T cell Acute Lymphoblastic Leukemia. Science 306 (5694) :269-271. doi: 10.1126/science.1102160.
3. Bhanushali AA, Babu S, Thangapandi VR, Pillai R, Chheda P, Das BR. (2010). Mutations in the HD and PEST domain of Notch-1 receptor in T-cell acute lymphoblastic leukemia: report of novel mutations from Indian population. Oncol Res. 19(2): 99-104.