You are here: Future directions
This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Future directions
As has been discussed in the Gene ontology page, Notch1 is involved in signaling events responsible for regulation of cellular proliferation and cell fate. Hence, the association of Notch1 with Acute T-cell Lymphoblastic Leukemia (T-ALL) is not surprising. In fact, many other cancer types as well as other diseases characterized by misregulation of cellular proliferation and/or cell fate decision also harbor mutations in Notch1. Some examples are multiple sclerosis, Alagille Syndrome, and Spondylocostal Dysostosis [1]. Consequently, a detailed understanding of Notch signaling pathway as well as its regulatory mechanisms is essential to shed light on potential treatments for these diseases. Although γ-secretase inhibitor, a drug used to inhibit activation of Notch1, was shown to have potential in controlling malignancies in model organisms and cell lines [2][3], there are many disadvantages associated with this drug. For instance, γ-secretase inhibitor is not specific to Notch1 and may inhibit other Notch proteins, thereby generating undesired consequences. Also, γ-secretase inhibitor may be toxic to the body if used for long-term treatment. Specifically, higher dosage of γ-secretase inhibitor has been reported to cause severe diarrhea in human patients [2].
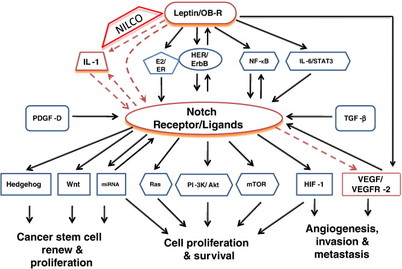
As with other signaling pathways, Notch signaling network involves complex crosstalks with multiple signaling pathways (see Figure 1). These crosstalks may be important for determining the ultimate outcome in response to a particular signaling molecule or environmental cues. Given the clinical relevance of Notch signaling, it will be interesting to dissect the interaction of Notch signaling pathway with other pathways, and understand how changes in a component in the network affects the expression or activities of other components in the network.
As with other signaling pathways, Notch signaling network involves complex crosstalks with multiple signaling pathways (see Figure 1). These crosstalks may be important for determining the ultimate outcome in response to a particular signaling molecule or environmental cues. Given the clinical relevance of Notch signaling, it will be interesting to dissect the interaction of Notch signaling pathway with other pathways, and understand how changes in a component in the network affects the expression or activities of other components in the network.
The lack of complete understanding on the network and mechanisms of Notch1 as well as the need for a safe and effective treatment of T-ALL prompted the proposal of the following experiments:
Experiment 1: Does age affect NOTCH1 expression level?
Hypothesis: NOTCH1 is expressed at a higher level in children and adults who are >70 years old, which contributes to their higher risk of developing ALL.
A larger proportion of acute lymphoblastic leukemia (ALL) occur in children, as shown in the pie chart on the home page. It has also been reported that the risk of developing ALL is higher not only in children, but also in adults who are over 70 years old. This seems to suggest that there might be an age factor that contribute to the observed distribution of ALL. Since Notch1 mutation is associated with ALL, it would be interesting to identify if NOTCH1 expression, as well as the expression of its target genes, is affected by age. The proposed method for this experiment is to carry out quantitative polymerase chain reaction (qPCR) analysis of the bone marrow, thymus and peripheral blood samples of mice from different age groups: juvenile, young adult, and geriatric (old) mice. The choice of these tissue types is based on their respective role in T-cell production, maturation, and circulation. If the hypothesis is correct and age does influence Notch1 expression and activation, the expected results would be the detection of alteration in the expression of NOTCH1 and its target genes, with a higher expression observed in juvenile and geriatric mice compared to the young adult mice. In fact, higher level of NOTCH1 expression has been reported in microarray analysis of umbilical cord blood samples of human infant compared to the bone marrow samples of human adults [ATLAS] (Figure 2). However, whether this is also true for older adults and whether this alteration also affects the expression of Notch1 target genes remain to be elucidated.
Hypothesis: NOTCH1 is expressed at a higher level in children and adults who are >70 years old, which contributes to their higher risk of developing ALL.
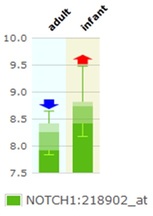
A larger proportion of acute lymphoblastic leukemia (ALL) occur in children, as shown in the pie chart on the home page. It has also been reported that the risk of developing ALL is higher not only in children, but also in adults who are over 70 years old. This seems to suggest that there might be an age factor that contribute to the observed distribution of ALL. Since Notch1 mutation is associated with ALL, it would be interesting to identify if NOTCH1 expression, as well as the expression of its target genes, is affected by age. The proposed method for this experiment is to carry out quantitative polymerase chain reaction (qPCR) analysis of the bone marrow, thymus and peripheral blood samples of mice from different age groups: juvenile, young adult, and geriatric (old) mice. The choice of these tissue types is based on their respective role in T-cell production, maturation, and circulation. If the hypothesis is correct and age does influence Notch1 expression and activation, the expected results would be the detection of alteration in the expression of NOTCH1 and its target genes, with a higher expression observed in juvenile and geriatric mice compared to the young adult mice. In fact, higher level of NOTCH1 expression has been reported in microarray analysis of umbilical cord blood samples of human infant compared to the bone marrow samples of human adults [ATLAS] (Figure 2). However, whether this is also true for older adults and whether this alteration also affects the expression of Notch1 target genes remain to be elucidated.
|
Figure 2: Higher expression of Notch1 in infants compared to adults in human. Figure adapted from ATLAS.
|
Experiment 2: What is the function of Domain of Unknown Function (DUF)?
Hypothesis: Domain of unknown function may interact with proteins that negatively regulate activated intracellular Notch1 stability.
As shown in Protein Domains page, Notch1 protein is reported to contain a domain of unknown function towards the end of the intracellular segment. Comparison of the domain location to the reported sites of T-ALL mutations by Weng, P. et al. [5], it seems likely that a group of T-ALL mutations fall into this Domain of Unknown Function (DUF). Since the intracellular domain of Notch1 is the part that acts as transcription factor upon Notch1 activation, and given that overactivation of Notch1 contributes to >50% of T-ALL cases, there is the possibility that the mutations reported in T-ALL affect the downregulation and/or turn-over of activated Notch1. Consequently, DUF might function as a site of negative regulation via interactions with proteins that confer post-translational modifications to this domain. To test whether this hypothesis is true, TAP-tag technique could be used to identify protein-protein interactions that occur specifically with this domain. The gene segment encoding the DUF domain can be fused to TAP tag and the resulting transgene could be transfected into T-lymphocyte cell line. After growing the cells to the appropriate confluence, the cells can be lysed and the tandem purification of proteins could be carried out, followed by mass spectrometry analysis to identify the interacting proteins. If the hypothesis is correct, the proteins isolated are expected to be those that can post-translationally modify DUF, such as ubiquitin ligase or kinases.
Experiment 3: Is there potential for combinatorial drug treatment?
Hypothesis: Combined targeting of Notch1 and β-catenin may enhance the efficacy of the treatment of Notch1-associated T-ALL.
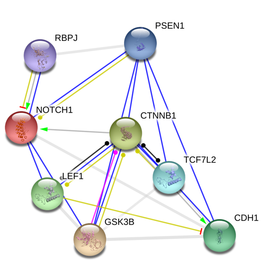
As discussed above, Notch1 signaling network is highly complex and it is very difficult to predict the outcome of a particular genetic lesion because the final outcome is likely dependent on components in other pathways. I am particularly interested to study the interaction between Wnt signaling and Notch signaling because components of the Wnt signaling pathway (CTNNB1, LEF1, GSK3B, TCF7L2) have been found to interact with Notch1 (Figure 3). The fact that these pathways interact with each other may suggest that the targeting of a single or a few components where the pathways converge may be sufficient to reverse or ameliorate the malignant phenotype. In addition, high-specificity treatments could be designed by targeting a common component in the interaction network along with another component specific to an observed malignancy. To test this hypothesis in the context of T-ALL, a lower dose of γ-secretase inhibitor could be combined with inhibitors of β-catenin (encoded by CTNNB1). The resulting drug combination could be tested on T-ALL murine model, and T-ALL disease progression as well as metastatic potential could be monitored. Mouse is a suitable model organism for this experiment because of the well-established T-ALL model available, and also because of the ability to evaluate whether combinatorial treatment is also effective against metastasis - an analysis that would not be possible with in vitro analysis. The advantage of combinatorial drug treatment is that lower doses of each drug could be prescribed because effects of the drugs are expected to be cumulative. With a lower dose, toxicity associated with a particular drug could be lowered (see above for a discussion of the side-effects of γ-secretase inhibitor). It might also be possible to tailor the dosage of each drug to achieve the treatment with highest specificity.
Figure 3: Protein-protein interactions of Notch1, after adding CTNNB1 as an input node. Figure from String9.0.
In summary, Notch1 is an important regulator of cellular proliferation and cell fate. It is therefore unsurprising that mutations in Notch1 are found in many types of cancer, making Notch1 one of the key targets in treatment design. However, despite many years of research, the complete network of Notch1 signaling and its mechanisms of action are not yet fully understood. Given the significance of Notch1 both in terms of pharmaceutical applications and basic biological research, more effort should be devoted to elucidate how Notch1 is regulated, how and what genes it regulates, and whether feedback loop exists in regulating Notch1 activation.
References:
1. Rampal, R., Luther, K.B., & Haltiwanger, R.S. (2007). Notch signaling in normal and disease states: possible therapies related to glycosylation. Curr Mol Med 7(4): 427-445. [PubMed]
2. Garber, K. (2007). Notch Emerges as New Cancer Drug Target. Journal of the National Cancer Institute. 99(17): 1284-1285.
3. Keersmaecker, K.D., Lahortiga, I., & Mentens, N. (2008). In vitro validation of γ-secretase inhibitors alone or in combination with other anti-cancer drugs for the treatment of T-cell acute lymphoblastic leukemia. Haematologica 93(4): 533-542.
4. Guo, S., Liu, M., & Gonzalez-Perez, R.R. (2011). Role of Notch and its oncogenic signaling crosstalk in breast cancer. Biochimica et Biophysica Acta - Reviews on Cancer 1815(2): 197-213
5. Weng, A.P., Ferrando, A.A., Lee, W., Morris IV, J.P., Silverman, L.B., Sanchez-Irizarry, C., Blacklow, S.C., Look, A.T., & Aster, J.C. (2004). Activating Mutations of NOTCH1 in Human T cell Acute Lymphoblastic Leukemia. Science 306 (5694) :269-271. doi: 10.1126/science.1102160.
1. Rampal, R., Luther, K.B., & Haltiwanger, R.S. (2007). Notch signaling in normal and disease states: possible therapies related to glycosylation. Curr Mol Med 7(4): 427-445. [PubMed]
2. Garber, K. (2007). Notch Emerges as New Cancer Drug Target. Journal of the National Cancer Institute. 99(17): 1284-1285.
3. Keersmaecker, K.D., Lahortiga, I., & Mentens, N. (2008). In vitro validation of γ-secretase inhibitors alone or in combination with other anti-cancer drugs for the treatment of T-cell acute lymphoblastic leukemia. Haematologica 93(4): 533-542.
4. Guo, S., Liu, M., & Gonzalez-Perez, R.R. (2011). Role of Notch and its oncogenic signaling crosstalk in breast cancer. Biochimica et Biophysica Acta - Reviews on Cancer 1815(2): 197-213
5. Weng, A.P., Ferrando, A.A., Lee, W., Morris IV, J.P., Silverman, L.B., Sanchez-Irizarry, C., Blacklow, S.C., Look, A.T., & Aster, J.C. (2004). Activating Mutations of NOTCH1 in Human T cell Acute Lymphoblastic Leukemia. Science 306 (5694) :269-271. doi: 10.1126/science.1102160.