This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Notch1 protein phylogeny
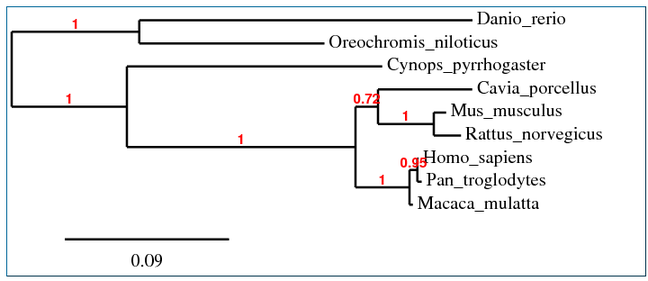
Phylogenetic tree 1:
Program: Phylogeny.fr (one-click mode, with Gblocks curation)
Algorithm: PhyML
Sequence alignment: MUSCLE
Program: Phylogeny.fr (one-click mode, with Gblocks curation)
Algorithm: PhyML
Sequence alignment: MUSCLE
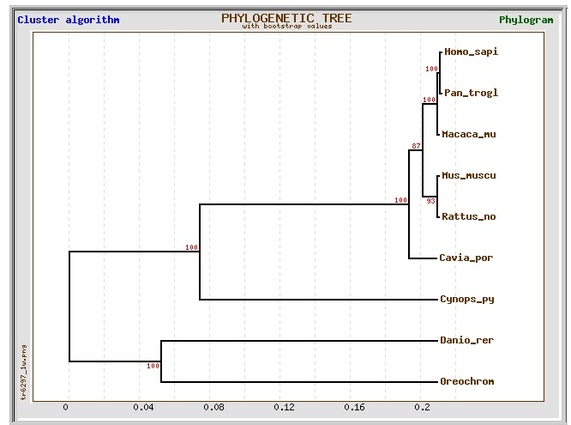
Phylogenetic tree 2:
Program: GeneBee
Algorithm: Cluster, with bootstrap values
Sequence alignment: ClustalW2
Program: GeneBee
Algorithm: Cluster, with bootstrap values
Sequence alignment: ClustalW2
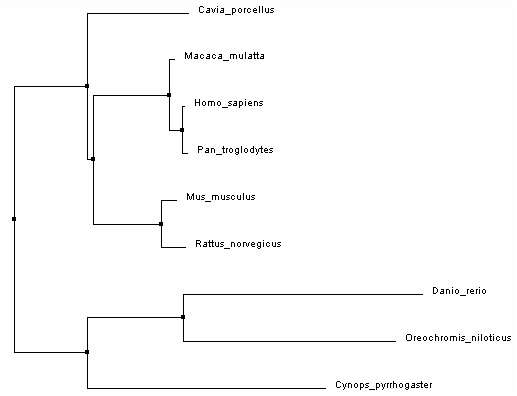
Phylogenetic tree 4:
Program: Jalview 2
Algorithm: Neighbor joining using percent identity
Alignment: MUSCLE
Program: Jalview 2
Algorithm: Neighbor joining using percent identity
Alignment: MUSCLE
Analysis:
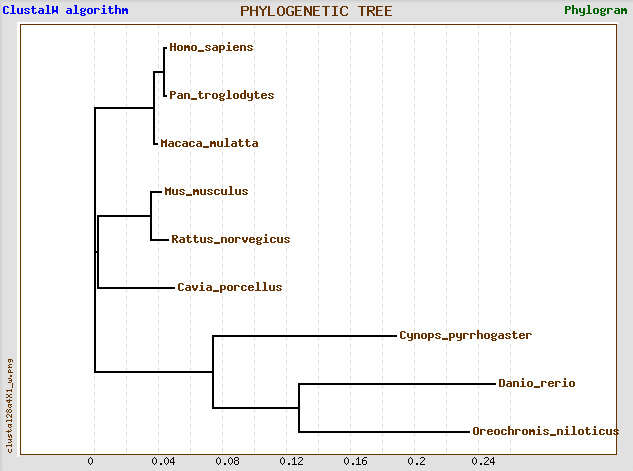
All of the phylogenetic trees above consist of 3 main branches, which will be referred to as branches A, B and C. Branch A includes Guinea pig (Cavia porcellus), mice (Mus musculus), rats (Rattus norvegicus), monkeys (Macaca mulatta), humans (Homo sapiens), and chimpanzees (Pan troglodytes); Branch B includes newt (Cynops pyrrhogaster); Branch C includes of zebrafish (Danio rerio) and nile tilapia (Oreochromis niloticus). In general, all 4 trees are similar except for the placement of Guinea pig within branch A. In trees 1 and 3, Guinea pigs, mice and rats share a common ancestor that diverged from the common ancestor of monkeys, humans, and chimpanzees. On the other hand, in trees 2 and 4, Guinea pigs diverged from the common ancestors of the other 5 organisms in branch A (rats, mice, monkeys, humans and chimpanzees). This is probably because different algorithms were used in different programs. Overall, the results were not surprising (humans, chimpanzees, and monkeys are more closely related to each other than to other organisms, so are mice and rats as well as tilapia and zebrafish).
References:
1. Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. (PubMed)
2. Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9. Epub 2008 Apr 19. (PubMed)*: joint first authors
3. Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, Mar 19;32(5):1792-7. (PubMed)
4. Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000, Apr;17(4):540-52. (PubMed)
5. Guindon S., Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, Oct;52(5):696-704. (PubMed)
6. Anisimova M., Gascuel O. Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 2006, Aug;55(4):539-52. (PubMed)
7. Chevenet F., Brun C., Banuls AL., Jacq B., Chisten R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics. 2006, Oct10;7:439. (PubMed)
8. Waterhouse, A.M., Procter, J.B., Martin, D.M.A, Clamp, M., Barton, G.J (2009), "Jalview version 2: A Multiple Sequence Alignment and Analysis Workbench," Bioinformatics 25 (9) 1189-1191 doi: 10.1093/bioinformatics/btp033