This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Microarray
Microarrays are "tool[s] for analyzing gene expression", and are usually made of "a small membrane or glass slide containing samples of many genes arranged in a regular pattern" [NCBI]. This tool allows the investigation of gene expression at the RNA level. Many experiments have been done to study the effects of NOTCH1 inhibition on various gene expression. This is not surprising given that NOTCH1 is long known to be involved in signaling events and transcriptional regulation. Shown below are some microarray data retrieved via GEO Profiles from the experiment carried out by Dohda, T. et al. (2007). The experiment analyzed total RNA isolated from T-ALL cell lines of Homo sapiens (human).
Dataset record number: GDS2794
Experiment description: "Analysis of T-cell acute lymphoblast leukemia (T-ALL) MOLT4 cells following gamma-secretase inhibitor (GSI) DAPT treatment. Gamma-secretase activity is required for Notch 1 receptor signaling. Results provide insight into the role of Notch signaling in T-ALL development."
Dataset record number: GDS2794
Experiment description: "Analysis of T-cell acute lymphoblast leukemia (T-ALL) MOLT4 cells following gamma-secretase inhibitor (GSI) DAPT treatment. Gamma-secretase activity is required for Notch 1 receptor signaling. Results provide insight into the role of Notch signaling in T-ALL development."
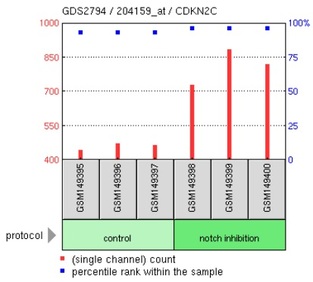
Figure 1: Gene expression profile for CDKN2C (Cyclin-dependent kinase inhibitor 2C) in response to notch inhibition via gamma-secretase inhibitor:
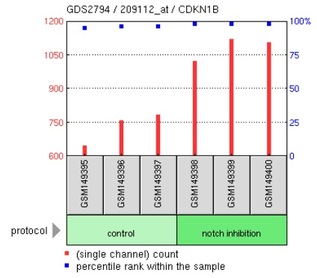
Figure 2: Expression of CDKN1B (Cyclin0dependent kinase inhibitor 1B) in response to notch inhibition:
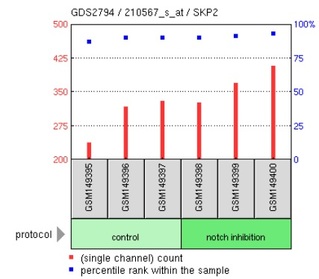
Figure 3: Expression of SKP2 (S-phase kinase-associated proteins) in response to notch inhibition.
Analysis:
Using the search terms "NOTCH1" and "T-ALL" in GEO Profiles, 3 distinct microarray experiments were returned. Part of the data from 1 of the experiments were shown above. In this particular experiment, 54675 expression profiles were reported. Since the focus of this project is on the association of Notch1 to T-ALL, 3 gene expression profiles that are related to T-ALL pathology were presented: CDKN2C, CDKN1B and SKP2. CDKN2C and CDKN1B are both inhibitors of cyclin-dependent kinases, which regulate progression through cell cycle. The microarray data revealed that inhibition of notch led to the increase in expression levels of CDKN2C and CDKN1B (also called P27Kip1) in human T-ALL cell lines, which could then inhibit the activity of cyclin-dependent kinases. This reduces the rate of abnormal cellular proliferation in the human T-ALL cell lines carrying activating mutation in NOTCH1. From the results, it seems possible that gamma-secretase inhibitors might be suitable for treating NOTCH1-associated T-ALL. Inhibition of notch also increases the expression of SKP2, which is a component of E3 ubiquitin-protein ligase complex that functions to regulate its target proteins via proteasomal degradation [UniProt]. The targets of SKP2 include proteins that are involved in G1/S transition stage of cell cycle. Hence, overexpression of SKP2 is likely to downregulate these protein targets and reduce the rate of cell proliferation. Overall, the microarray data showed that NOTCH1 is an important transcriptional regulator of many genes, some of which are related to cellular proliferation. Therefore, it is not surprising that mutations in NOTCH1 contribute to T-ALL, which is characterized by overproliferation of T-cell blasts.
References:
1. GEO Profiles
2. Dohda T, Maljukova A, Liu L, Heyman M, Grandér D, Brodin D, Sangfelt O, Lendahl U. Notch signaling induces SKP2 expression and promotes reduction of p27Kip1 in T-cell acute lymphoblastic leukemia cell lines. Exp Cell Res. 2007 Aug 15;313(14):3141-52. Epub 2007 May 5. PubMed PMID: 17560996.
3. UniProt